Scientific Keyword Discovery Hub Raphaelepsis Explaining Biological Research Queries
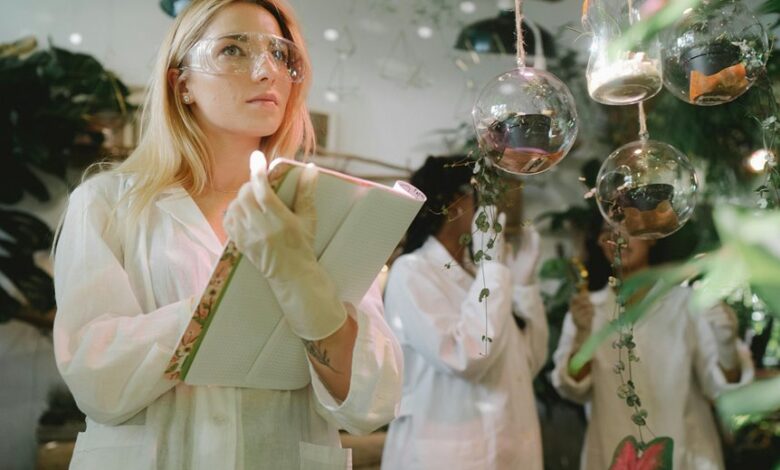

Raphaelepsis functions as a bridge between biology and data science, translating keywords into interoperable vocabularies. It identifies terms from literature, databases, and experiments, disambiguating synonyms and aligning contexts across ontologies. The approach yields transparent workflow steps, scalable validation, and reproducible search strategies. This framework clarifies intent and reveals latent connections, guiding researchers toward robust hypothesis generation. The next question arises: how will this mapping perform across diverse datasets and disciplines?
What Is Scientific Keyword Discovery in Biology?
Scientific keyword discovery in biology refers to the systematic identification of terms and phrases that best represent biological concepts, data, and research questions within a corpus of literature, databases, or experimental results. The process involves disambiguating terms and aligning cross disciplinary synonyms to enhance searchability, comparison, and reproducibility across studies, databases, and analytical frameworks. Clear terminology supports transparent scientific communication and efficient knowledge integration.
How Raphaelepsis Maps Terms Across Disciplines
Raphaelepsis maps terms across disciplines by linking concepts that share semantic targets despite differing vocabularies. It employs cross disciplinary term mapping to reveal equivalences between domain-specific lexicons, enabling interoperable search and analysis. The process emphasizes keyword interoperability, aligning synonyms, hierarchies, and contextual usage. Through structured mappings, researchers access related concepts across fields, supporting transparent, scalable integration without conflating distinct disciplinary nuances.
A Practical Workflow for Building/Validating Keywords
A practical workflow for building and validating keywords operationalizes the cross-disciplinary mappings established previously by detailing step-by-step methods, criteria, and checks. The process emphasizes emerging term normalization to maintain consistency across datasets and disciplines, while enforcing cross ontology linkage to ensure semantic interoperability, traceability, and versioning. This disciplined approach supports transparent refinement, reproducibility, and scalable keyword validation across research domains.
Real-World Queries: From Datasets to Discovery Signals
How do real-world queries transform raw datasets into actionable discovery signals? Real-world queries synthesize heterogeneous data, normalize variability, and extract robust features for interpretation. The process emphasizes reproducibility, transparent metrics, and scalable pipelines. Techniques comparison reveals performance trade-offs, while cross disciplinary mapping aligns domains to reveal novel insights. This approach supports freedom-driven inquiry by enabling targeted, evidence-based hypothesis generation and rapid validation.
Conclusion
Raphaelepsis provides a precise, cross-ontology framework for translating biology terms into interoperable vocabularies, aligning synonyms and contexts to reveal searchable intent. By mapping keywords across disciplines, it clarifies queries, fosters reproducible searches, and supports validated workflows. Its transparent steps enable scalable hypothesis generation and rapid signaling from datasets to discovery. An anachronistic flourish—a medieval scriptorium annotating CRISPR—illustrates how structured keywords hold timeless power to illuminate complex biology in any era.